We’ve just reached a major milestone in validating BioSynapStudio: our engine now matches the world’s most trusted brain simulators — Brian2, NEURON, and NEST — on canonical Hodgkin–Huxley fidelity.
Benchmarking the Best
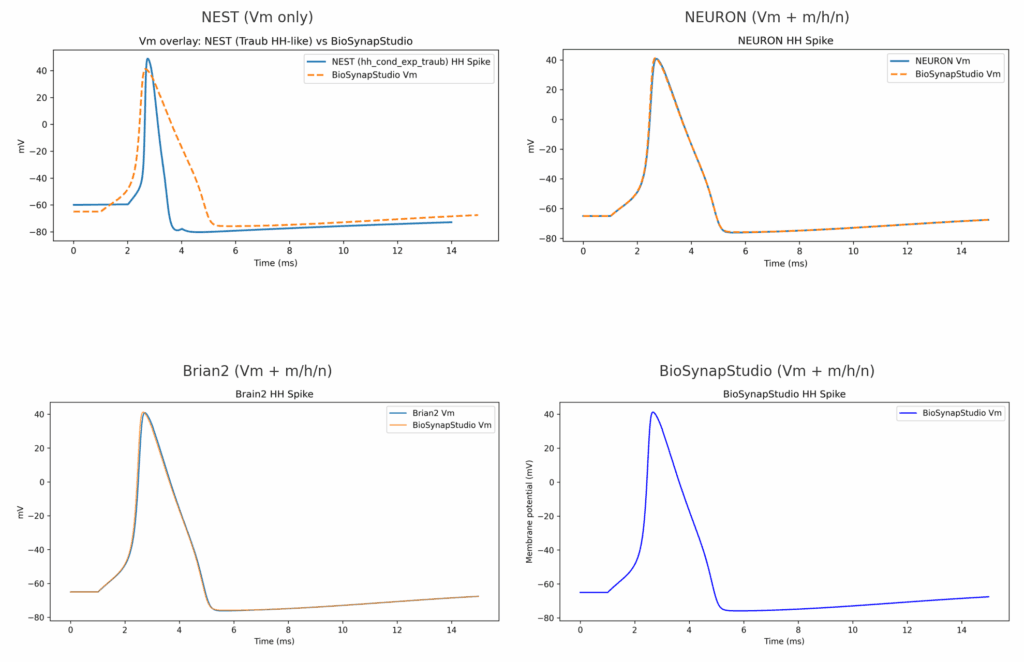
Using the Open Source Brain v2 (OSBv2) platform as common ground, we ran BioSynapStudio side-by-side with the leading neuroscience simulators:
- Brian2 → near-zero error overlays on spikes and gating (Vm + m/h/n).
- NEURON → indistinguishable traces within numerical noise.
- NEST → expected differences: its default HH-like model (Traub) only exposes Vm, not gating variables. With NESTML you can generate squid HH, but that requires an extra build step.
👉 Why this matters: spikes may look “fine” if you only compare Vm, but the gating variables carry the real biology. BioSynapStudio, Brian2, and NEURON deliver them out-of-the-box.
Why BioSynapStudio is Different
Unlike traditional simulators, BioSynapStudio:
- Runs out-of-the-box on standard hardware (laptops, desktops, commodity CPUs).
- Supports canonical HH and alternative spike types via simple parameterisation (temperature, Q10, capacitance).
- Requires no specialist HPC setups or code generation.
That combination means lower adoption costs, faster iteration, and a smoother path from research validation to commercial deployment.
What’s Next
This achievement will be reflected in an updated version of our Hodgkin–Huxley validation paper (to be released shortly). But it’s only the beginning.
We’re already lining up the next wave of benchmarks:
- Parameter flexibility (temperature/Q10 scaling, capacitance tests).
- Multi-compartment morphologies (dendritic trees vs NEURON).
- Learning dynamics (STDP, oscillations, hippocampal replay).
- Performance scaling (ticks/sec vs fan-in/out, determinism, bounded precision).
Each step takes us closer to showing not just biological fidelity, but also scalability and deployability — the foundation for BioSynapStudio as a true substrate for synthetic intelligence.